Protein structures
BioJava provides an API to maintain local installations of the PDB, load and manipulate structures, perform standard analysis such as sequence and structure alignments and visualize them in 3D.
The project is hosted, developed, and maintained on GitHub.
GitHub project
BioJava provides an API to maintain local installations of the PDB, load and manipulate structures, perform standard analysis such as sequence and structure alignments and visualize them in 3D.
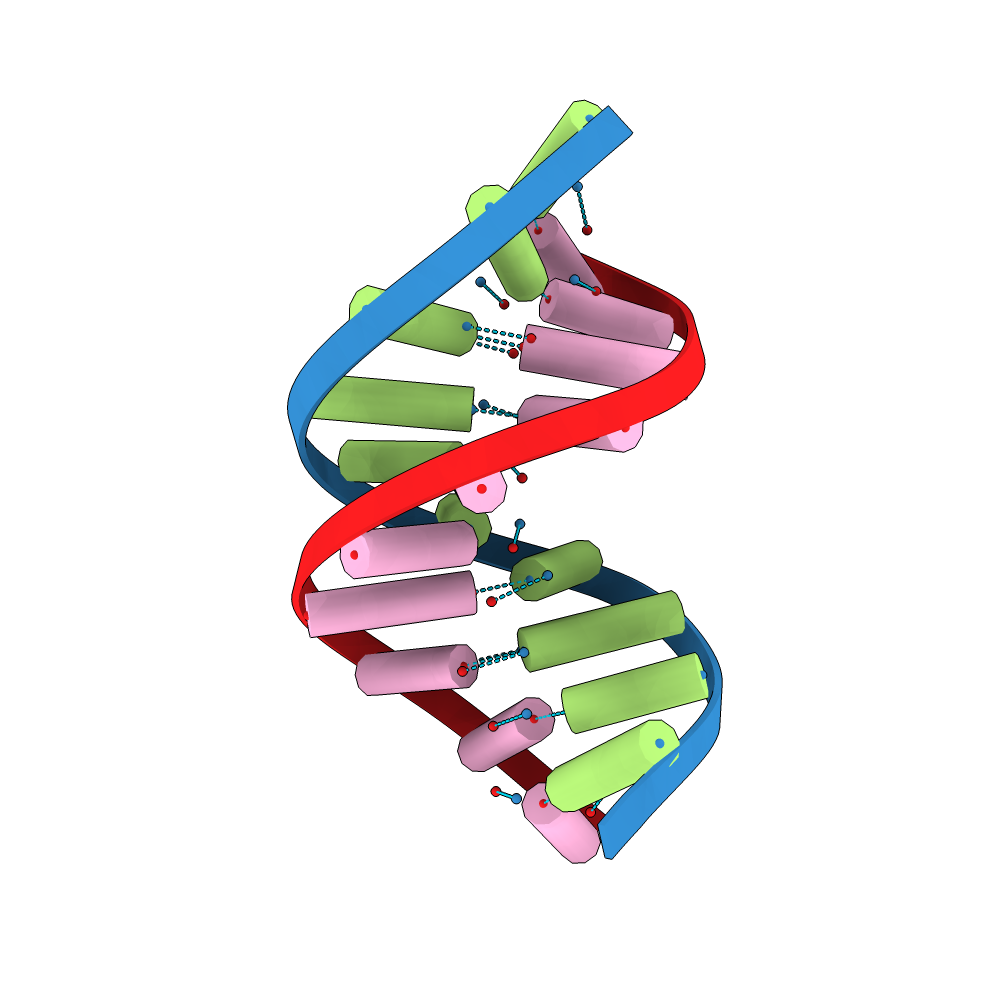

BioJava supports reading and writing popular sequence file formats, translating DNA sequences into proteins and other common bioinformatics routines.
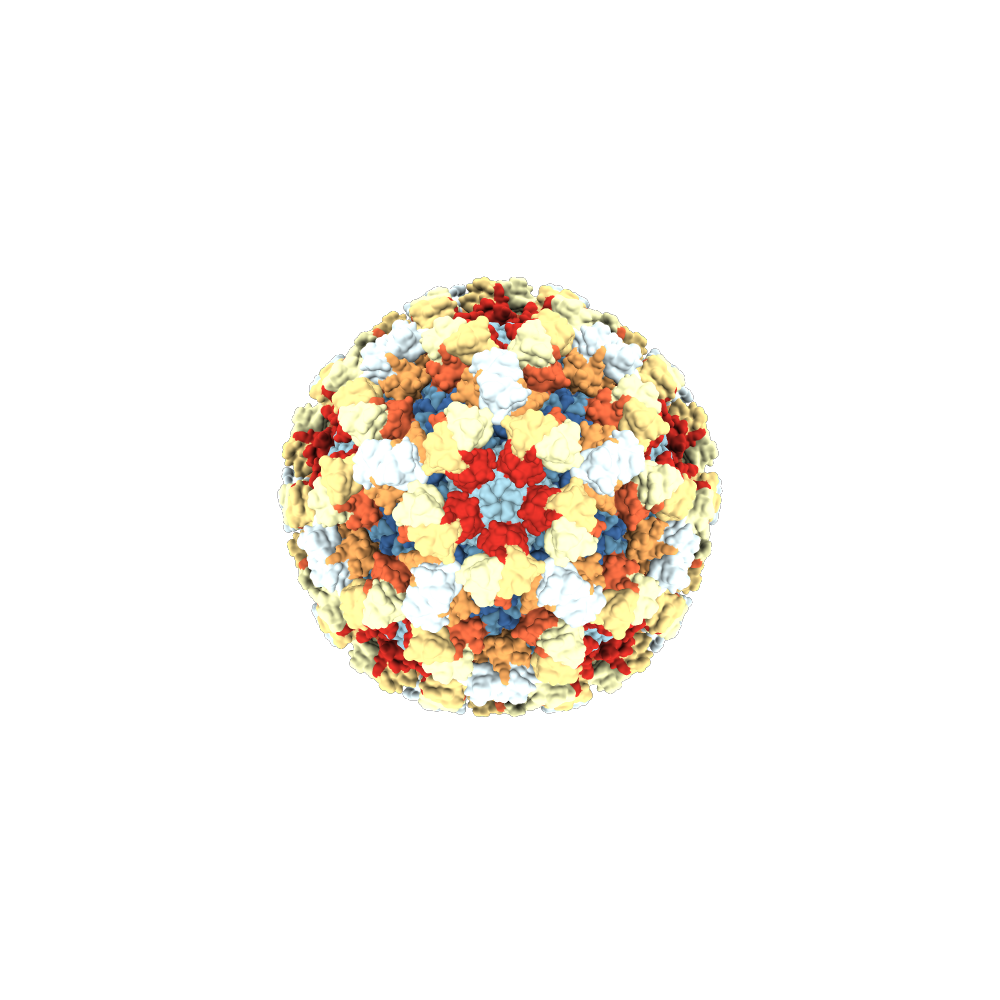

BioJava 5: A community driven open-source bioinformatics library
Aleix Lafita, Spencer Bliven, Andreas Prlić, Dmytro Guzenko, Peter W. Rose, Anthony Bradley, Paolo Pavan, Douglas Myers-Turnbull, Yana Valasatava, Michael Heuer, Matt Larson, Stephen K. Burley, & Jose M. Duarte
PLOS Computational Biology (2019) 15 (2):e1006791.

Questions, comments, and communications regarding BioJava can be done either on GitHub or through the BioJava mailing list. Subscribe to stay up to date!
The tutorial offers an introduction into some of the features provided by BioJava.
TutorialThe Cookbook provides simple coding recipes to common bioinformatics problems using BioJava.
CookbookThe Javadocs for the current BioJava 7.1.4 release.
Javadoc APIBioJava is available from Maven Central.
Maven CentralAll pages from the legacy wiki site have been migrated to markdown pages
View wikiThe Javadocs for BioJava legacy 1.x versions.
View 1.9.5 APIHere a couple of pointers to get you started